Yang, Q.L., Xie, Y., Qiao, K., Lim, J.Y.S., and S. Wu. (2025). Modern biology of extrachromosomal DNA: A decade-long voyage of discovery. Cell Research 35, 11–22. (PubMed)
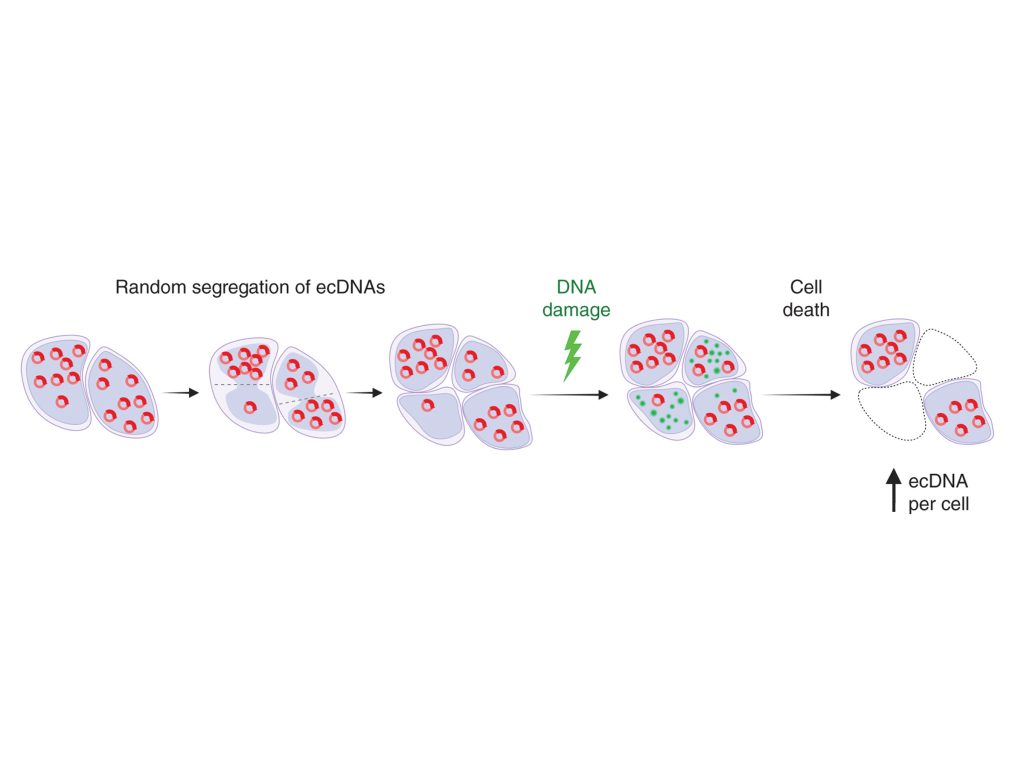
Lin, M.S., Jo, S.Y., Luebeck, J., Chang, H.Y., Wu, S., Mischel, P.S., Bafna, V. (2024). Transcriptional immune suppression and up-regulation of double-stranded DNA damage and repair repertoires in ecDNA-containing tumors. Elife 12 :RP88895. (PubMed)
Pal Choudhuri, S., Girard, L., Lim, J.Y.S., Wise, J.F., Freitas, B., Yang, D., Wong, E., Hamilton, S., Chien, V.D., Kim, Y.J., Gilbreath, C., Zhong, J., Phat, S., Myers, D.T., Christensen, C.L., Mazloom-Farsibaf, H., Stanzione, M., Wong, K.K., Hung, Y.P., Farago, A.F., Meador, C.B., Dyson, N.J., Lawrence, M.S.,# Wu S.,# and B.J. Drapkin.# (2024). Acquired Cross-resistance in Small Cell Lung Cancer due to Extrachromosomal DNA Amplification of MYC paralogs. Cancer Discovery 14, 804-827. (PubMed)
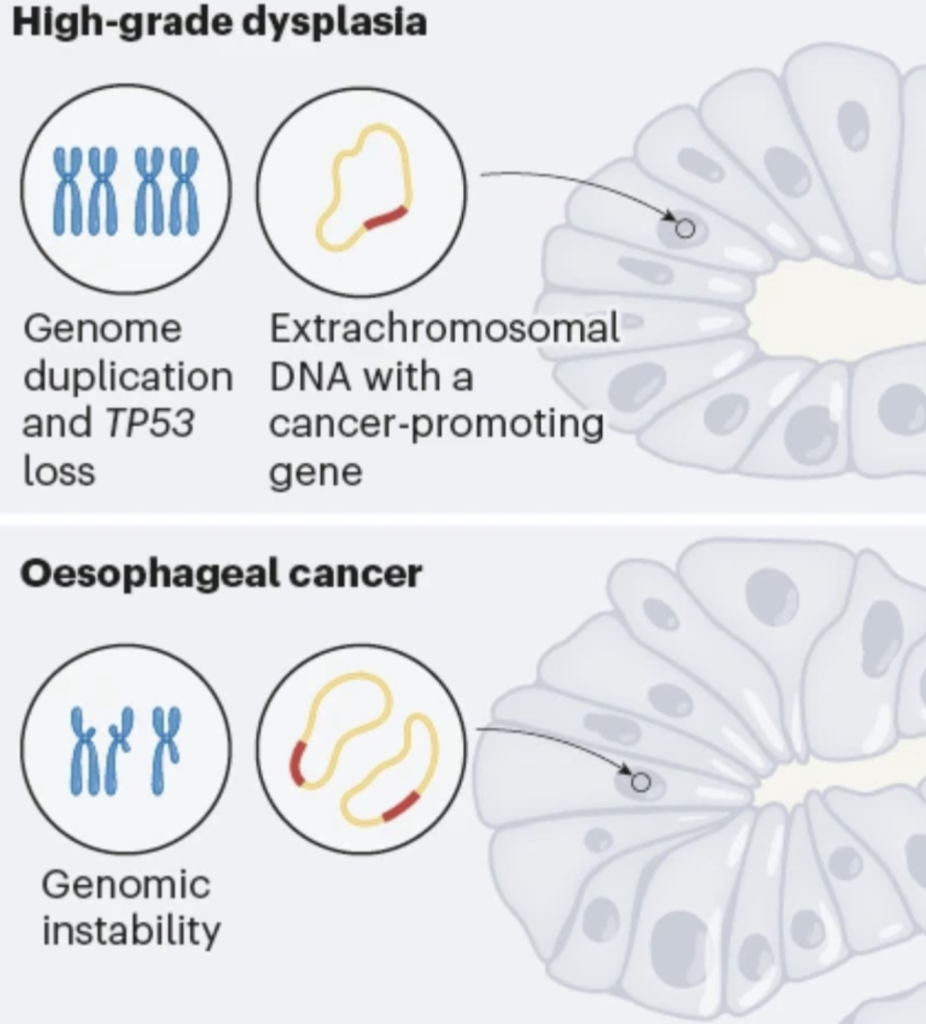
Luebeck, J., Ng, A.W.T., Galipeau, P.C., Li, X., Sanchez, C.A., Katz-Summercorn, A., Kim, H., Jammula, S., He, Y., Lippman, S.M., Verhaak, R.G.W., Maley, C.C., Alexandrov, L.B., Reid, B.J., Fitzgerald, R.C., # Paulson, T.G.,# Chang, H.Y.,# Wu, S.,# Bafna, V.,# and P.S. Mischel.# (2023). Extrachromosomal DNA in the cancerous transformation of Barrett’s esophagus. Nature 616, 798-805. (PubMed)
Chen, P.B., Fiaux, P.C., Zhang, K., Li, B., Kubo, N., Jiang, S., Hu, R., Rooholfada, E., Wu, S., Wang, M., Wang, W., McVicker, G., Mischel, P.S., and B. Ren. (2022). Systematic discovery and functional dissection of enhancers needed for cancer cell fitness and proliferation. Cell Reports 41, 111630. (PubMed)
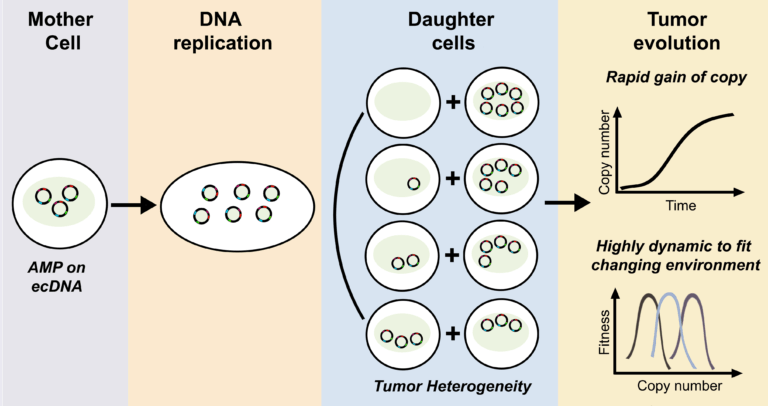
Lange, J.T., Rose, J.C., Chen, C.Y., Pichugin, Y., Xie, L., Tang, J., Hung, K.L., Yost, K.E., Shi, Q., Erb, M.L., Rajkumar, U., Wu, S., Taschner-Mandl, S., Bernkopf, M., Swanton, C., Liu, Z., Huang, W., Chang, H.Y., Bafna, V., Henssen, A.G., Werner, B., and P.S. Mischel. (2022). The evolutionary dynamics of extrachromosomal DNA in human cancers. Nature Genetics 54, 1527-1533. (PubMed)
Hung, K.L., Yost, K.E., Xie, L., Shi, Q., Helmsauer, K., Luebeck, J., Schöpflin, R., Lange, J.T., Chamorro González, R., Weiser, N.E., Chen, C., Valieva, M.E., Wong, I.T., Wu, S., Dehkordi, S.R., Duffy, C.V., Kraft, K., Tang, J., Belk, J.A., Rose, J.C., Corces, M.R., Granja, J.M., Li, R., Rajkumar, U., Friedlein, J., Bagchi, A., Satpathy, A.T., Tjian, R., Mundlos, S., Bafna, V., Henssen, A.G., Mischel, P.S., Liu, Z., and H.Y. Chang. (2022). ecDNA hubs drive cooperative intermolecular oncogene expression. Nature 600, 731-736. (PubMed)
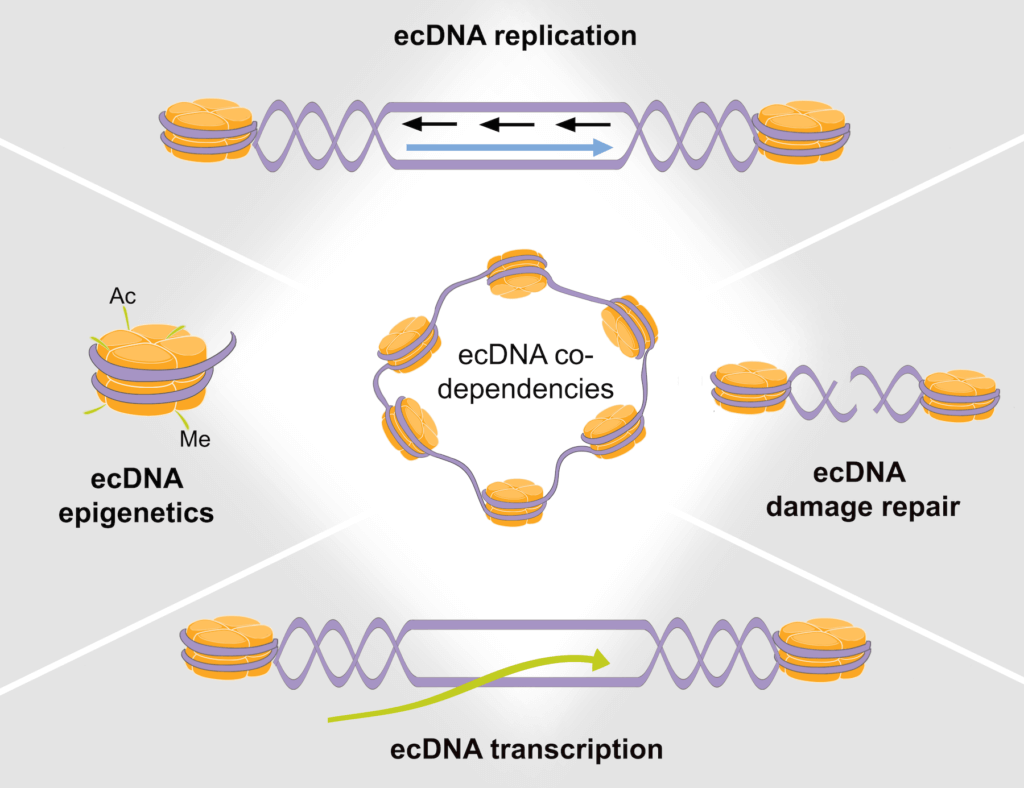
Wu, S., Bafna, V., Chang, H. Y., and P.S. Mischel. (2022). Extrachromosomal DNA: An Emerging Hallmark in Human Cancer. Annual Review of Pathology: Mechanisms of Disease 17, 367-386. (PubMed)
Wu, S., Bafna, V., and P.S. Mischel. (2021). Extrachromosomal DNA (EcDNA) in Cancer Pathogenesis. Current Opinion in Genetics & Development 66, 78–82. (PubMed)
Kim, H., Nguyen, N.-P., Turner, K., Wu, S., Gujar, A. D., Luebeck, J., Liu, J., Deshpande, V., Rajkumar, U., Namburi, S., Amin, S. B., Yi, E., Menghi, F., Schulte, J. H., Henssen, A. G., Chang, H. Y., Beck, C. R., Mischel, P. S., Bafna, V., and R.G.W. Verhaak. (2020). Extrachromosomal DNA Is Associated with Oncogene Amplification and Poor Outcome across Multiple Cancers. Nature Genetics 52, 891–897. (PubMed)
Cai, J., Liu, W., Wong, C. W., Zhu, W., Lin, Y., Hu, J., Xu, W., Zhang, J., Sander, M., Wang, Z., Dan, J., Zhang, J., Liu, Y., Guo, L., Qin, Z., Liu, X., Liu, Y., Yan, G., Wu, S., and J. Liang. (2020). Zinc-Finger Antiviral Protein Acts as a Tumor Suppressor in Colorectal Cancer. Oncogene 39, 5995–6008. (PubMed)
Bi, J., Chowdhry, S., Wu, S., Zhang, W., Masui, K., and P.S. Mischel. (2020). Altered Cellular Metabolism in Gliomas — an Emerging Landscape of Actionable Co-Dependency Targets. Nature Reviews Cancer 20, 57–70. (PubMed)
Wu, S., Turner, K. M., Nguyen, N., Raviram, R., Erb, M., Santini, J., Luebeck, J., Rajkumar, U., Diao, Y., Li, B., Zhang, W., Jameson, N., Corces, M. R., Granja, J. M., Chen, X., Coruh, C., Abnousi, A., Houston, J., Ye, Z., Hu, R., Yu, M., Kim, H., Law, J. A., Verhaak, R. G. W., Hu, M., Furnari, F. B., Chang, H. Y., Ren, B., Bafna, V., and P.S. Mischel. (2019). Circular EcDNA Promotes Accessible Chromatin and High Oncogene Expression. Nature 575, 699–703. (PubMed)
#Co-corresponding author