Guo, J.*, Liang, R.*, Chung, A.*, Li, Z.*, Li, B., Chen, E., Li, L., Wang, J., Hsieh, M.H., Fang, I.X., Kroger, B., Wang, Y., Zhu, M., Ren, X., Mannino, G., Jia, Y., Wei, Y., Moore, S., Siegwart, D.J., Chung, S.S., Wang, Z., Sharma, T., Komjeti, S., Han, Y., Gopal, P., Xiao, G., Wang, T., and H. Zhu. (2025). The origin of hepatocellular carcinoma depends on metabolic zonation. Science 391:eadv7129. (PubMed)
Zhu, M., Wang, Y., Lu, T., Guo, J., Li, L., Hsieh, M.H., Gopal, P., Han, Y., Fujiwara, N., Wallace, D.P., Yu, A.S.L., Fang, X., Ransom, C., Verschleisser, S., Hsiehchen, D., Hoshida, Y., Singal, A.G., Yopp, A., Wang, T., and H. Zhu. PKD1 mutant clones within cirrhotic livers inhibit steatohepatitis without promoting cancer. (2024). Cell Metabolism 36, 1711-25. (PubMed)
Wang, Z., Zhu, S., Jia, Y., Wang, Y., Kubota, N., Fujiwara, N., Gordillo, R., Lewis, C., Zhu, M., Sharma, T., Li, L., Zeng, Q., Lin, Y.H., Hsieh, M.H., Gopal, P., Wang, T., Hoare, M., Campbell, P., Hoshida, Y., and H. Zhu. (2023). Positive selection of somatically mutated clones identifies adaptive pathways in metabolic liver disease. Cell 186, 1968-1984. (PubMed)
Lin, Y.H., Wei, Y., Zeng, Q., Wang, Y., Pagani, C.A., Li, L., Zhu, M., Wang, Z., Hsieh, M.H., Corbitt, N., Zhang, Y., Sharma, T., Wang, T., and H. Zhu. (2023). IGFBP2 expressing midlobular hepatocytes preferentially contribute to liver homeostasis and regeneration. Cell Stem Cell 30, 665-676. (PubMed)
Jia, Y., Li, L., Lin, Y.H., Gopal, P., Shen, S., Zhou, K., Yu, X., Sharma, T., Zhang, Y., Siegwart, D., Ready, J., and H. Zhu. (2022). In vivo CRISPR screening identifies BAZ2 chromatin remodelers as druggable regulators of mammalian liver regeneration. Cell Stem Cell 3, 372-385.(PubMed)
Wei, Y., Wang, Y.G., Jia, Y., Li, L., Yoon, J., Zhang, S., Wang, Z., Zhang, Y., Zhu, M., Sharma, T., Lin, Y.H., Hsieh, M.H., Albrecht, J.H., Le, P.T., Rosen, C.J., Wang, T., and H. Zhu. (2021). Liver homeostasis is maintained by midlobular zone 2 hepatocytes. Science 371:eabb162. (PubMed)
Zhu, M., Lu, T., Jia, Y., Luo, X., Gopal, P., Li, L., Odewole, M., Renteria, V., Singal, A.G., Jang, Y., Ge, K., Wang, S.C., Sorouri, M., Parekh, J.R., MacConmara, M.P., Yopp, A.C., Wang, T., and H. Zhu. (2019). Somatic Mutations Increase Hepatic Clonal Fitness and Regeneration in Chronic Liver Disease. Cell 177, 608-621. (PubMed)
Zhang, S., Zhou, K., Luo, X., Li, L., Tu, H.C., Sehgal, A., Nguyen, L.H., Zhang, Yu., Gopal, P., Tarlow, B., Siegwart, D.J., and H. Zhu. (2018). The polyploid state plays a tumor-suppressive role in the liver. Developmental Cell 44, 447-459. (PubMed)
Sun, X.,* Wang, S.C.,* Luo, X., Jia, Y., Li, L., Gopal, P., Zhu, M., Nassour, I., Chuang, J.C., Maples, T., Celen, C., Nguyen, L.H., Wu, L., Fu, S., Li, W., Hui, L., Tian, F., Ji, Y., Zhang, S., Sorouri, M., Hwang, T.H., Letzig, L., James, L., Yopp, A., Singal, A., and H. Zhu. (2017). Arid1a has context-dependent oncogenic and tumor suppressor functions in liver cancer. Cancer Cell 32, 574-589. (PubMed)
Sun, X., Chuang, J.C., Kanchwala, M., Wu, L., Celen, C., Li, L., Liang, H., Zhang, S., Maples, T., Nguyen, L.H., Wang, S.C., Signer, R.A., Sorouri, M., Nassour, I., Liu, X., Xu, J., Wu, M., Zhao, Y., Kuo, Y.C., Wang, Z., Xing, C., and H. Zhu. (2016). Suppression of the SWI/SNF Component Arid1a Promotes Mammalian Regeneration. Cell Stem Cell 18, 456–466. (PubMed)
Nguyen, L.H.,* Robinton, D.A.,* Seligson, M.T.,* Wu, L., Li, L., Rakheja, D., Comerford, S.A., Ramezani, S., Sun, X., Parikh, M.S., Yang, E.H., Powers, J.T., Shinoda, G., Shah, S.P., Hammer, R.E., Daley, G.Q.,# and H. Zhu.# (2014). Lin28b is sufficient to drive liver cancer and necessary for its maintenance in murine models. Cancer Cell 26, 248–261. (PubMed)
Shyh-Chang, N.,* Zhu, H.,* de Soysa, N.T., Shinoda, G., Seligson, M.T., Tsanov, K., Nguyen, L., Asara, J.M., Cantley, L.C. and Daley, G.Q. (2013). Lin28 enhances tissue repair by reprogramming cellular metabolism. Cell 155: 778-92. (PubMed)
Zhu, H.,* Shyh-Chang, N.,* Segrè, A.V., Shinoda, G., Shah, S.P., Einhorn, W.S., Takeuchi, A., Engreitz, J.M., Hagan, J.P., Kharas, M.G., Urbach, A., Thornton, J.E., Triboulet, R.,nGregory, R.I.; DIAGRAM Consortium; MAGIC Investigators, Altshuler, D., and Daley, G.Q. (2011). The Lin28/let-7 axis regulates glucose metabolism. Cell 147: 81-94. (PubMed)
*Contributed equally; #Co-corresponding author



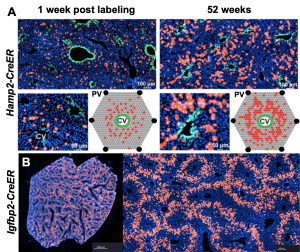 The cellular basis of liver regeneration has been highly contentious. We generated 12 CreER strains to perform systematic, side-by-side comparisons of hepatocytes in various zones, including populations previously implicated as stem cells (
The cellular basis of liver regeneration has been highly contentious. We generated 12 CreER strains to perform systematic, side-by-side comparisons of hepatocytes in various zones, including populations previously implicated as stem cells (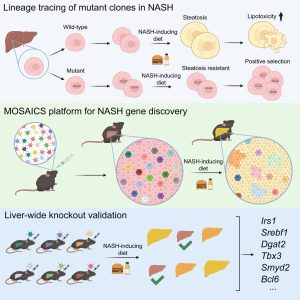 Somatic mosaicism is a new frontier for human genetics and represents an untapped source of disease genes. While somatic mutations can now be identified using deep sequencing, the functional analysis of such mutations remains challenging and will require new approaches. We are currently using in vivo screening and lineage tracing as well as CRISPR technologies to rapidly model the phenotypic impact of somatic mutations. Just as germline sequencing has identified genes that have transformed fields, the discovery of genes that are recurrently mutated in chronically injured tissues but not in cancer has the potential to transform our understanding of regeneration and wound healing.
Somatic mosaicism is a new frontier for human genetics and represents an untapped source of disease genes. While somatic mutations can now be identified using deep sequencing, the functional analysis of such mutations remains challenging and will require new approaches. We are currently using in vivo screening and lineage tracing as well as CRISPR technologies to rapidly model the phenotypic impact of somatic mutations. Just as germline sequencing has identified genes that have transformed fields, the discovery of genes that are recurrently mutated in chronically injured tissues but not in cancer has the potential to transform our understanding of regeneration and wound healing.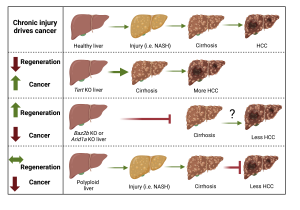 It is widely assumed that cancer risk increases with regenerative capacity, but we suspect the relationship is more complicated. In mammals, chronic organ damage in the skin, lung, intestine, and liver is strongly associated with cancer, but it is possible that the potent regenerative abilities of these organs serve to preserve tissue integrity, reduce inflammation, and resist transformation in the context of recurrent injury.
It is widely assumed that cancer risk increases with regenerative capacity, but we suspect the relationship is more complicated. In mammals, chronic organ damage in the skin, lung, intestine, and liver is strongly associated with cancer, but it is possible that the potent regenerative abilities of these organs serve to preserve tissue integrity, reduce inflammation, and resist transformation in the context of recurrent injury.











































